Bioequivalence
Bioequivalence vs
Non-inferiority vs Inferiority
When
conducting a clinical trial, a hypothesis of interest needs to be identified and
defined. Various statistical techniques can be used to address the chosen
hypothesis, depending on the type of comparison that is being made. The most
common comparison is between the effects of a new treatment compared to the
current treatment. That said, the specific comparison of interest will shape
the hypothesis and in turn, the statistical method required.
There
are three possible hypothesis that can be formulated when comparing two
treatments:
1.
The new treatment is at least as effective
as the current treatment (non-inferiority)
2.
The new treatment is strictly more
effective than the current treatment (inferiority)
3.
The new treatment is equivalent to the
current treatment (bioequivalence)
While
these three possible hypothesis appear to be similar, they are slightly
different, and thus require slightly different statistical tests.
Non-inferiority
trails are used to test whether a minimum level of efficacy has been achieved
in the new treatment. These trials are
designed to show that the new treatment is no less effective than the existing
treatment.
Inferiority/superiority
trials are used to test whether the new treatment is strictly more effective
than the current treatment. These trials are designed to detect a difference
between treatments.
Bioequivalence
trials are used to test whether a new treatment is the same, within an
acceptable range, to the current treatment. These trials are designed to
confirm the absence of a meaningful difference between treatments.
Before
we define the hypothesis’ and their corresponding statistical test, it
important to identify the type of outcome variable that is of interest.
Outcome Variable
Type
2.1
Binary Outcome

![]()
![]()
![]()
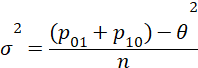
Unpaired
Non-inferiority:
![]() vs
vs ![]()
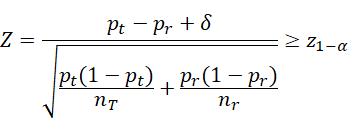
Inferiority:
![]() vs
vs ![]()
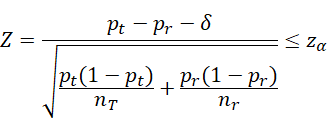
Bioequivalence:
![]() vs
vs ![]()
 and
and 
Code Example in SAS (non-inferiority)
/* Binary outcome */
/* Unpaired */
data table_unpair;
/* Data */
p1=0.4203;
p2=0.4638;
n1=69;n2=69;
/* Calculation */
theta=p1-p2;
var=(p1*(1-p1)/n1)+(p2*(1-p2)/n2);
se=sqrt(var);
delta=0.1; /* Pick a value of delta */
zright=(theta+delta)/se;
/* z-score for Non-inferiority */
zleft=(theta-delta)/se; /* z-score
for Inferiority */
pright=1-cdf("normal",zright); /* p-value for Non-inferiority */
pleft=cdf("normal",zleft); /* p-value for
Inferiority */
pboth=pright+pleft;
/* p-value for Bioequivalence */
run;
/* Paired */
data table;
/* Data */
p1=0.4203; *p1=proportion of 1 in protocol 1;
p2=0.4638; *p2=proportion of 1 in protocol 2;
p10=0;
p01=0.0435;
n=69;
/* Calculation */
theta=p1-p2;
var=(p10+p01-theta*theta)/n;
se=sqrt(var);
delta=0.1; /* Pick a value of delta */
zright=(theta+delta)/se;
*two-tailed test; /* z-score for Non-inferiority */
zleft=(theta-delta)/se;
*two-tailed test; /* z-score for Inferiority */
pright=1-cdf("normal",zright); /* p-value for
Non-inferiority */
pleft=cdf("normal",zleft); /* p-value for
Inferiority */
pboth=pright+pleft;
/* p-value for Bioequivalence */
run;
/* Continuous outcome */
/* Unpaired */
data Process;
input _STAT_ $4. @6 Process $3. @10 Quality Yield Waste;
cards;
N New 27 27 27
MEAN New 34.6667 50.0000 11.9889
STD
New 1.8605 0.8321 2.2548
N Old 27 27 27
MEAN Old 19.9530 40.1481 10.3556
STD
Old 0.8077 0.7698 2.1445
;
/* Non-inferiority */
proc ttest data=Process sides=U
h0=-0.1 /*delta*/ ;
class Process;
var Waste;
run;
/* Inferiority */
proc ttest data=Process sides=L
h0=0.1 /*delta*/ ;
class Process;
var Waste;
run;
/* Bioequivalence: p-value of Non-inferiority
+ p-value of Inferiority */
/* Paired */
DATA WEIGHT;
INPUT WBEFORE WAFTER;
DATALINES;
200 185
175 154
188 176
198 193
197 198
310 275
245 224
202 188
;
RUN;
/* Non-inferiority */
proc ttest data=WEIGHT sides=U
h0=-0.1 /*delta*/ ;
Paired WBEFORE * WAFTER;
run;
/* Inferiority */
proc ttest data=WEIGHT sides=L
h0=0.1 /*delta*/ ;
Paired WBEFORE * WAFTER;
run;
/* Bioequivalence: p-value of Non-inferiority + p-value of Inferiority */
Code Example in R (non-inferiority)
install.packages("tidyverse")
library(tidyverse)
# Binary
# Unpaired
p1=0.4203
p2=0.4638
theta=p1-p2
n1=69;n2=69
var=(p1*(1-p1)/n1)+(p2*(1-p2)/n2)
se=sqrt(var)
delta=0.1
zright=(theta+delta)/se
zleft=(theta-delta)/se
pright=1-pnorm(zright)
pleft=pnorm(zleft)
print(paste("Non-inferiority:
", "z-score:", zright,
"p-value:", pright))
print(paste("Inferiority:
", "z-score:", zleft,
"p-value:", pleft))
print(paste("Bioequivalence:
", "p-value:", pright+pleft))
# Paired
p1=0.4203
p2=0.4638
p10=0
p01=0.0435
theta=p1-p2
n=69
var=(p10+p01-theta*theta)/n
se=sqrt(var)
delta=0.1
zright=(theta+delta)/se
zleft=(theta-delta)/se
pright=1-pnorm(zright)
pleft=pnorm(zleft)
print(paste("Non-inferiority:
", "z-score:", zright,
"p-value:", pright))
print(paste("Inferiority:
", "z-score:", zleft,
"p-value:", pleft))
print(paste("Bioequivalence: ",
"p-value:", pright+pleft))
# Continuous
# Unpaired
# Data
process = data.frame(
process = c(rep("NEW",
27), rep("OLD", 27)),
quality = c(rnorm(27, 34.6667, 1.8605), rnorm(27, 19.9530, 0.8077)),
yield = c(rnorm(27, 50, 0.8321), rnorm(27, 40.1481, 0.7698)),
waste = c(rnorm(27, 11.9889, 2.2548), rnorm(27, 10.3556, 2.1445))
)
process_new = filter(process,
process == "NEW")
process_old = filter(process,
process == "OLD")
## Set a delta value
delta = 0.1
## Non-inferiority
t.test(process_new$waste, process_old$waste, mu = -delta, alternative =
"greater")
## Inferiority
t.test(process_new$waste, process_old$waste, mu = delta, alternative =
"less")
## Bioequivalence
p1 = t.test(process_new$waste, process_old$waste, mu = delta, alternative =
"less")$p.value
p2 = t.test(process_new$waste, process_old$waste, mu = -delta, alternative =
"greater")$p.value
print(paste("Bioequivalence
p-value: ", p1+p2))
# Paired
# Data
weight = data.frame(
wbefore
= c(200,175,188,198,197,310,245,202),
wafter = c(185,154,176,193,198,275,224,188)
)
## Non-inferiority
t.test(weight$wbefore, weight$wafter, paired = T, mu = -delta, alternative =
"greater")
## Inferiority
t.test(weight$wbefore, weight$wafter, paired = T, mu = delta, alternative =
"less")
## Bioequivalence
p1 = t.test(weight$wbefore, weight$wafter, paired = T, mu = delta, alternative =
"less")$p.value
p2 = t.test(weight$wbefore, weight$wafter, paired = T, mu = -delta, alternative =
"greater")$p.value
print(paste("Bioequivalence p-value: ", p1+p2))
References
2.
Althunian, T. A., de Boer, A., Groenwold, R. H. H., & Klungel, O.
H. (2017). Defining the noninferiority margin and analysing noninferiority: An overview. British Journal
of Clinical Pharmacology
3.
Committee for Proprietary Medicinal
Products. (2001). Points to consider on switching between superiority and
non-inferiority. British Journal of Clinical Pharmacology
4.
Tsong, Y., Zhang, J., & Levenson, M. (2007). Choice of δ
noninferiority margin and dependency of the noninferiority trials.
Journal of Biopharmaceutical Statistics
5.
Wang, B., Wang, H., Tu, X. M., & Feng, C. (2017). Comparisons of
Superiority, Non-inferiority, and Equivalence Trials. Shanghai Archives of
Psychiatry